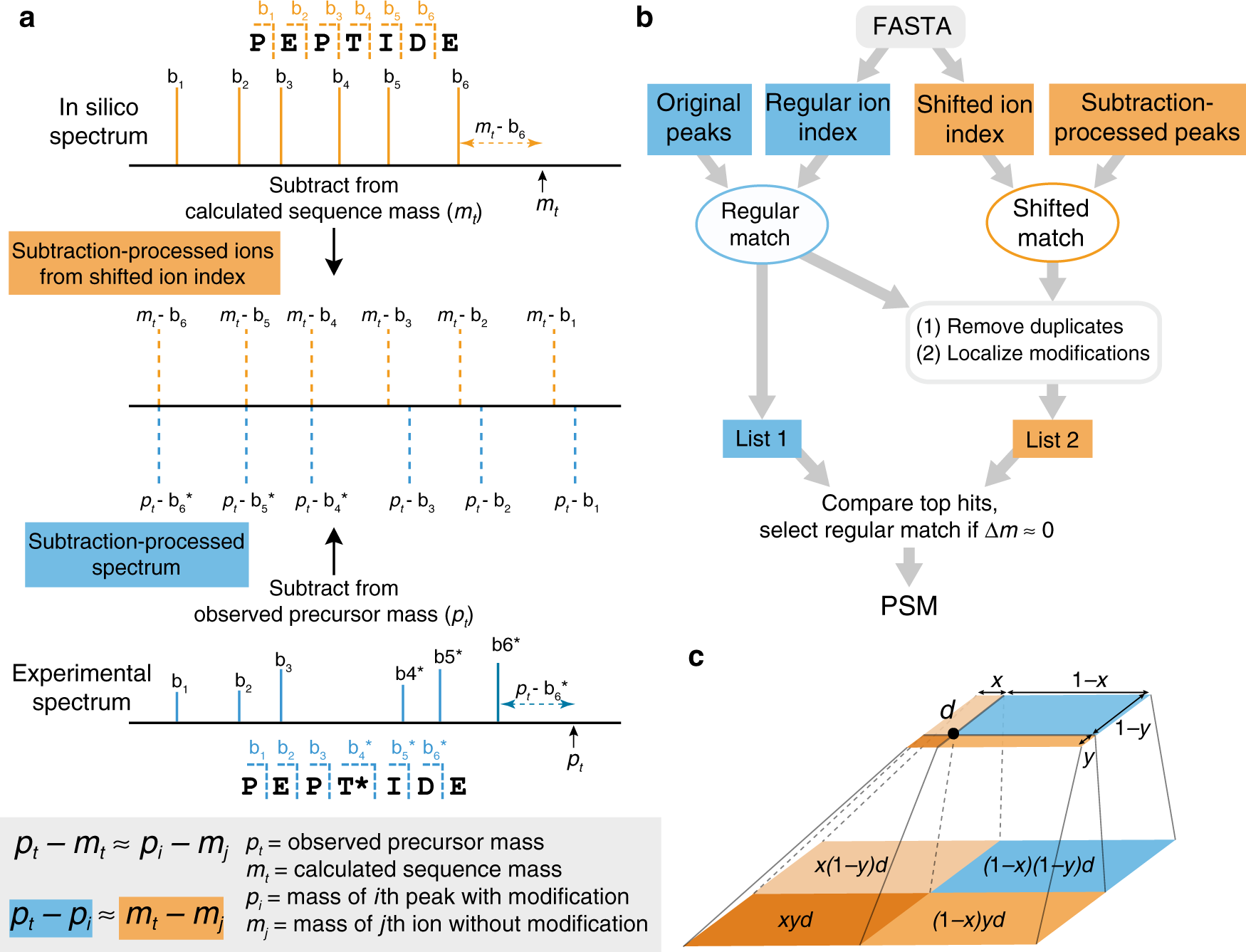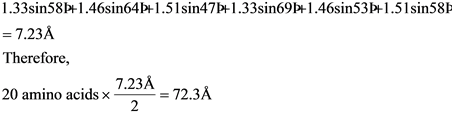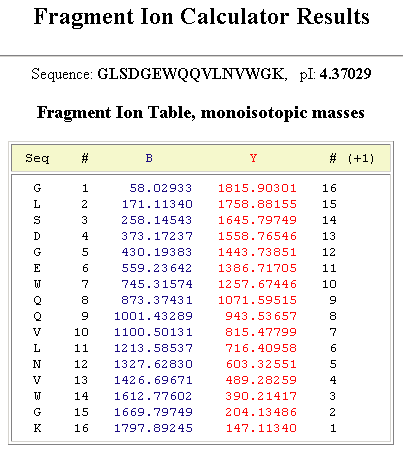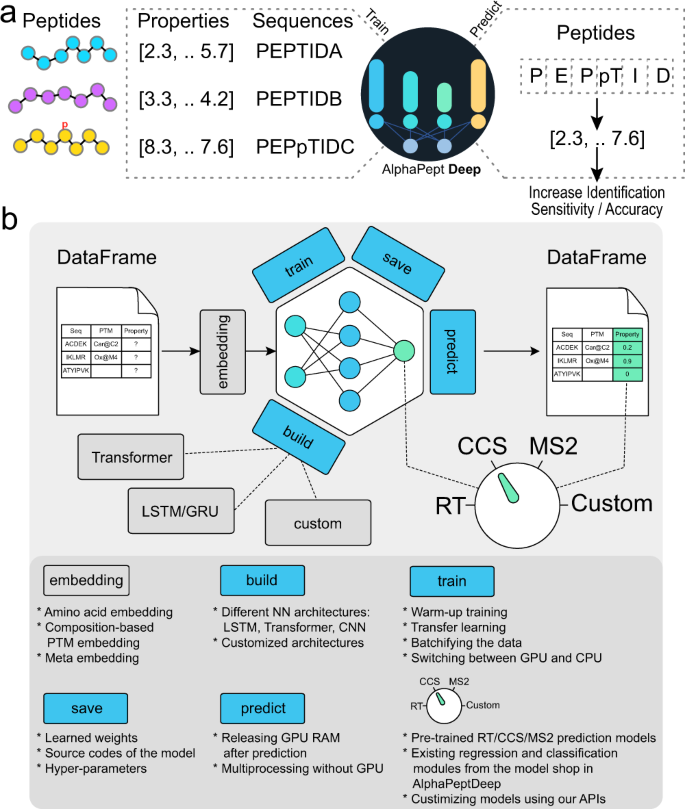
AlphaPeptDeep: a modular deep learning framework to predict peptide properties for proteomics | Nature Communications
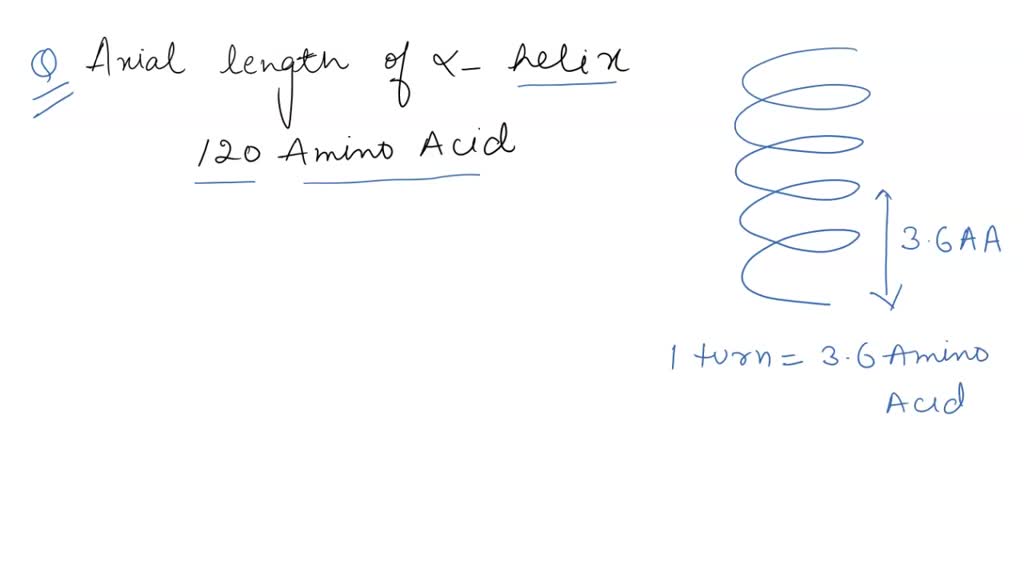
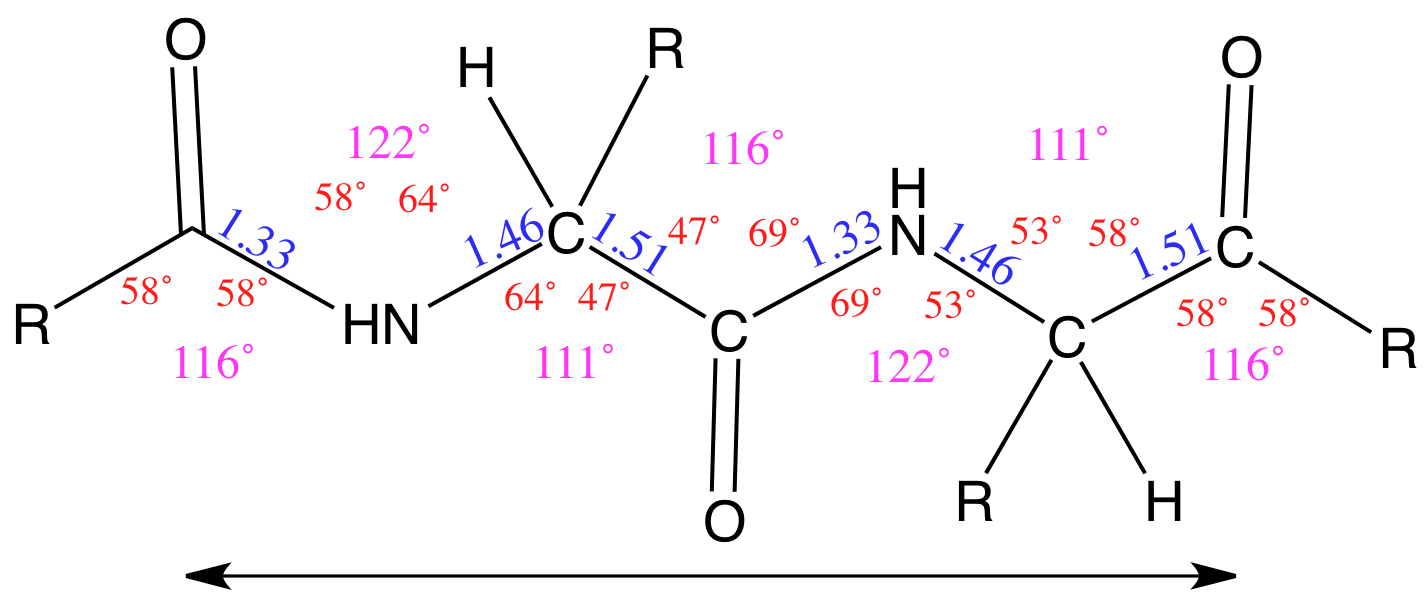
SOLVED: Calculate the axial length of an alpha helix that is of 120 amino acids long? How long would the polypeptide be if it were fully extended?
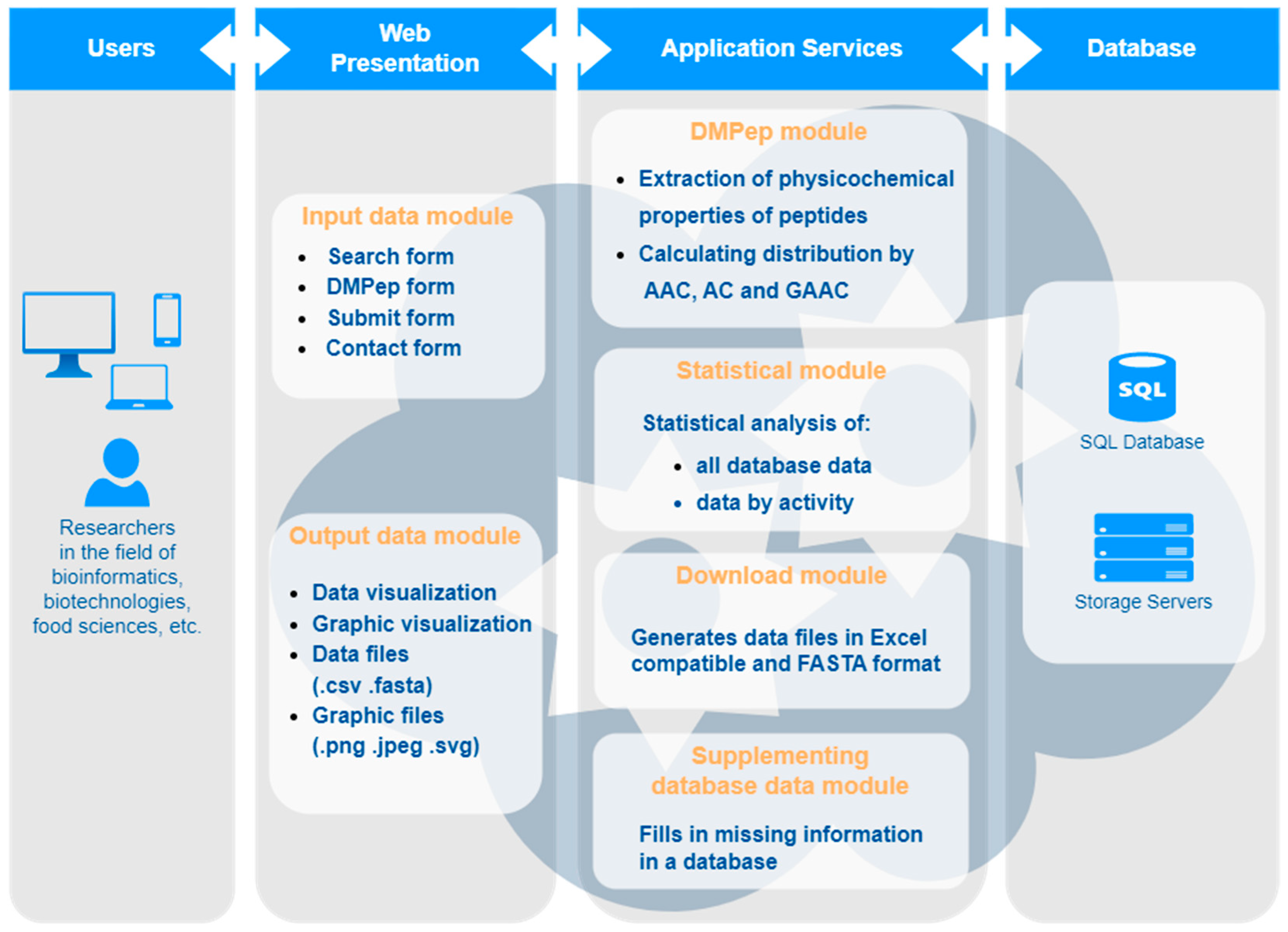
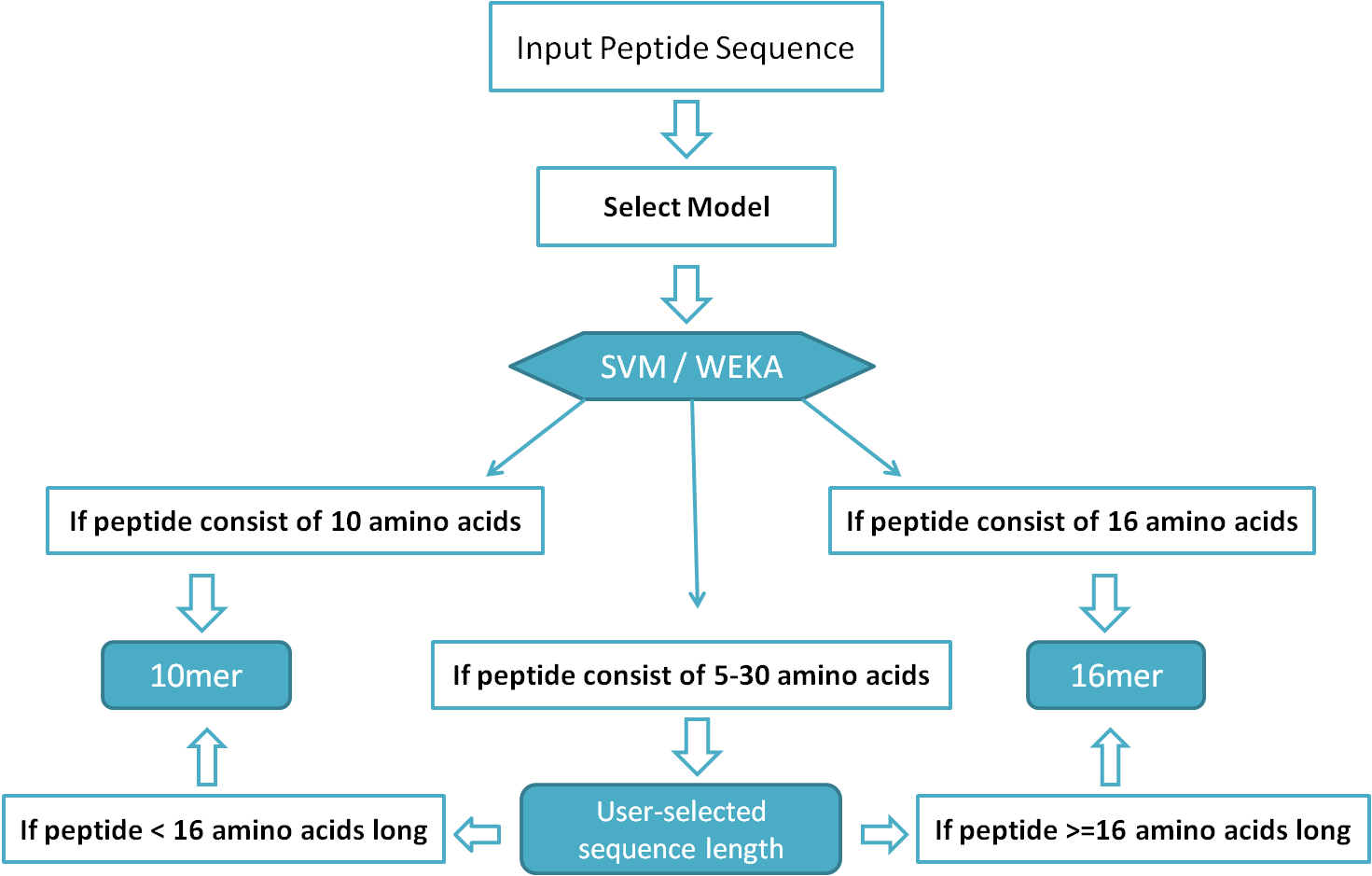
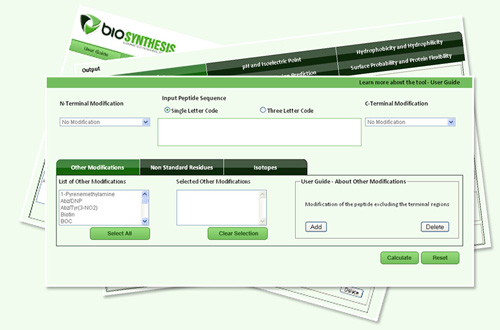
Applied Sciences | Free Full-Text | PepLab Platform: Database and Software Tools for Analysis of Food-Derived Bioactive Peptides

Sequence-Specific Model for Predicting Peptide Collision Cross Section Values in Proteomic Ion Mobility Spectrometry | Journal of Proteome Research

PeptiDesCalculator: Software for computation of peptide descriptors. Definition, implementation and case studies for 9 bioactivity endpoints - Barigye - 2021 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library
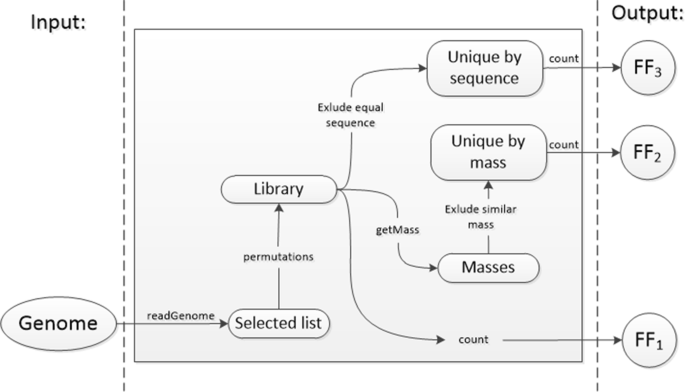
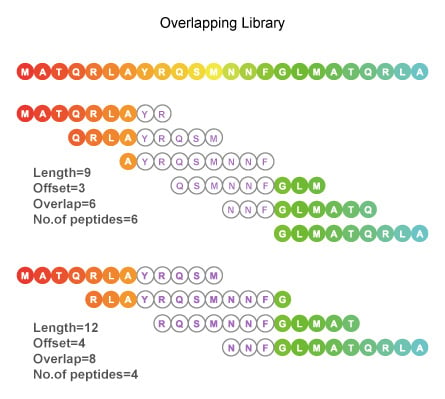
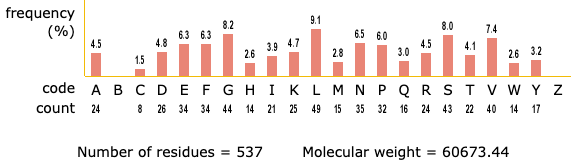
Algorithm-supported, mass and sequence diversity-oriented random peptide library design | Journal of Cheminformatics | Full Text

IJMS | Free Full-Text | Computational Evolution of Beta-2-Microglobulin Binding Peptides for Nanopatterned Surface Sensors